IRIS-HEP COVID-19 related activities
Like many scientific research and research computing teams, IRIS-HEP is organizing to contribute its software and computing capabilities and team in support of activities to meet the challenge of the COVID-19 pandemic. Some examples of our activities include:
Current Activities
-
An article, entitled “Stuck at home, physicists pivot to combat COVID-19” was published in Physics Today highlighting the COVID-19 efforts of the physics community. The article includes work by Peter Elmer, Henry Schreiner, Jim Pivarski and David Lange with the Princeton Open Ventilation Monitor project and Savannah Thais’ efforts with Science Responds.
-
UNL Research Professor Derek Weitzel and others from the Open Science Grid team are working with resource providers to enable and prioritize COVID-19 related workflows. New providers are joining the OSG to increase capacity for COVID-19 research. Usage of the OSG by COVID-19 workflows is accounted on GRACC, OSG’s accounting service. Over 38 institutions have contributed to OSG’s COVID-19 effort. Multiple COVID-19 workflows are utilizing the OSG, with more starting every week.
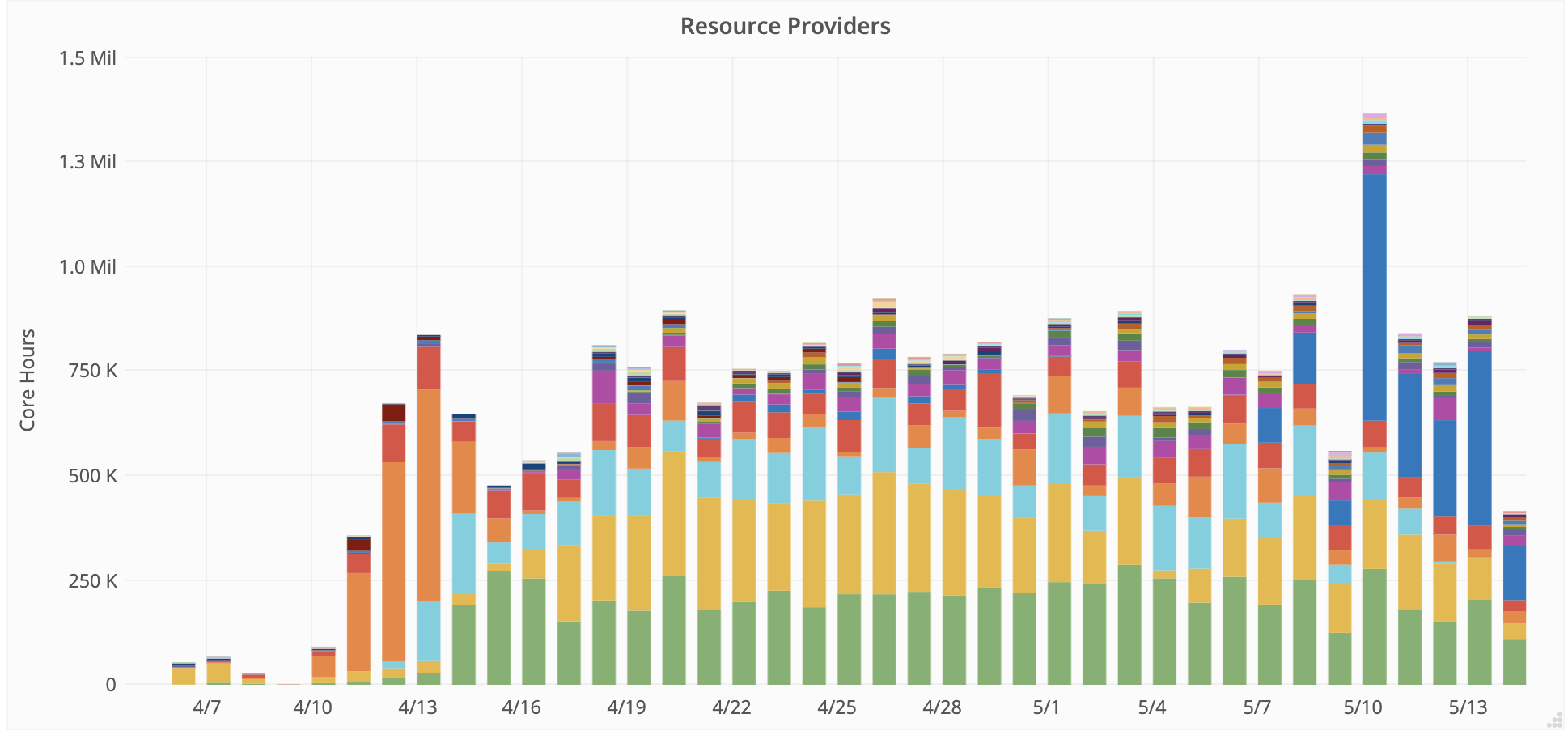
OSG COVID-19 Computing Hours contributed by Site. Over 38 Institutions have contributed to OSG’s COVID-19 research efforts.
-
University of Illinois at Urbana-Champaign postdoc Matthew Feickert has worked with the CERN COVID-19 Compute Response team to containerize the Folding@Home software in custom GPU-enabled Docker images to efficiently run Folding@Home on OSG sites with GPUs with donated resource time from ATLAS.
-
The Molecular Sciences Software Institute (MolSSI) is working with BioExcel on a COVID-19 Computational Molecular Data Exchange to help curate and provide the possibility of community review of the rapidly increasing number of COVID-19 related computational molecular simulation datasets. IRIS-HEP is discussing with them ideas for how HEP infrastructure such as Zenodo and the HEP data caching and distribution infrastructures might be something they can leverage as their curated data volumes increase.
-
Princeton postdoc Savannah Thais has organized a session at the APS April Meeting entitled “Response of Physics to the Coronavirus Pandemic”.
-
Princeton researchers Peter Elmer, Henry Schreiner, David Lange and Jim Pivarski are contributing the software for the Princeton Open Ventilation Monitor system, a patient pressure and airflow monitoring system for ventilators. The system allows up to 20 patients to be monitored remotely at a central station by a clinician or nurse in a COVID-19 field hospital, with relevant alarms. This outreach activity is being done in collaboration with the local multi-hospital healthcare system and a number of other Princeton Physics, Mechanical Engineering and Neuroscience faculty. The system includes data analysis algorithms, visualization and data acquisition from the sensor system (written in Python). The following image shows the nurse monitoring station GUI with simulated time series data for airflow, lung pressure and tidal volume transferred to the lungs. (Click for larger image.) This outreach activity has also spawned a separately funded research activity (PHY-2031509) by NSF. A preprint article summarizing this work is available: “Inexpensive multi-patient respiratory monitoring system for helmet ventilation during COVID-19 pandemic”
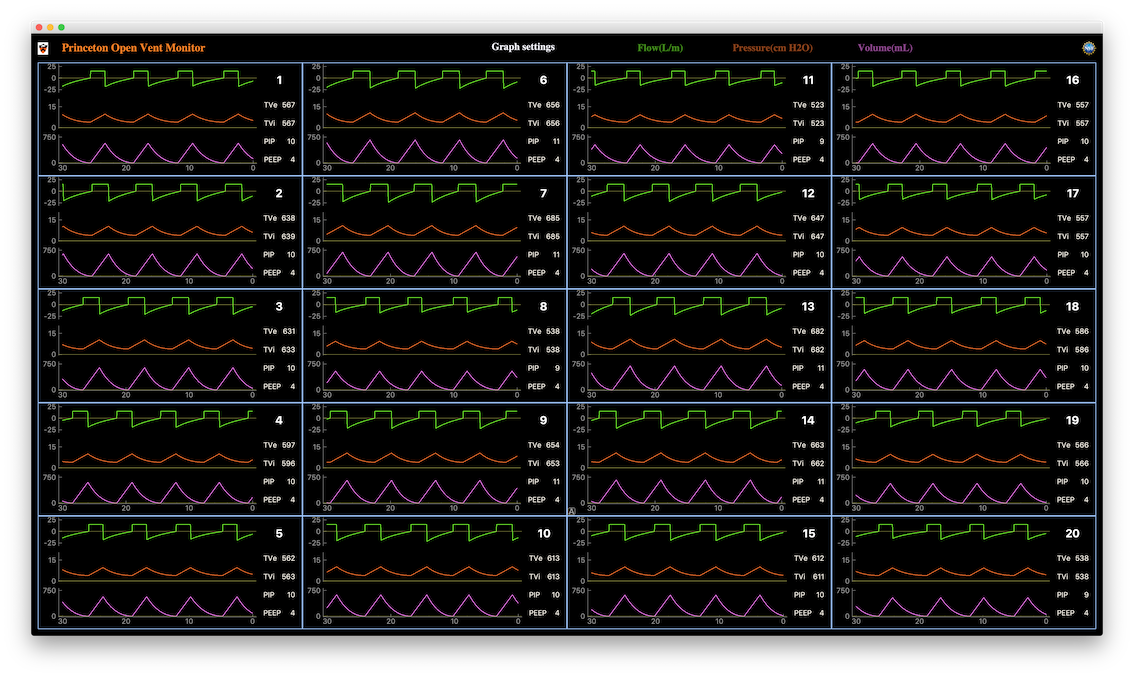
-
A number of IRIS-HEP researchers were involved in setting up the Science Responds to make connections that enable (non-medical) researchers to contribute to understanding and combating the global COVID-19 problem and/or its economic and societal impacts. This website provides resources and information for such researchers. An associated Slack workspace and Zoom meetings allow researchers to interact. We have also recently begun organizing talks and discussions with non-Physics researchers. Some of these are being recorded and made available in a Youtube channel.
-
University of Illinois Research Programmer, Ben Galewsky put data wrangling and website development skills to use to migrate the Champaign Urbana Health District’s spreadsheet of food distribution volunteers into a new NationBuilder site he set up. It allows staff to build community and put eager volunteers into action.
Earlier Relevant Activities
- Earlier IRIS-HEP work by the NYU Professor Kyle Cranmer and collaborators (“Efficient Probabilistic Inference in the Quest for Physics Beyond the Standard Model”, arXiv:1807.07706 ) established PPX, a probabilistic programming protocol. It has previously been extended for epidemiological studies such as “Hijacking Malaria Simulators with Probabilistic Programming”, arXiv:1905.12432 and is now also being applied to COVID19 (see “Planning as inference in epidemiological dynamics models” by Warrington, A., Naderiparizi, S., Weilbach, C., Masrani, V., Harvey, W., Scibior, A., Beronov, B., & Nasseri, A. (2020) arXiv:2003.13221 ).